
Solved Questions 1 Using The Amino Acid Numbers Provided Chegg Enhanced with ai, our expert help has broken down your problem into an easy to learn solution you can count on. see answer. question: use the table below to determine which four amino acids will be added next to the growing polypeptide? answer in order from left to right. use the table below to determine which four amino acids will be added. The mrna sequence "ucag" corresponds to the amino acid sequence serine glycine (ser gly) based on the information in the provided table. in the genetic code, each three base sequence, or codon, in mrna corresponds to a specific amino acid or a stop signal.

Solved Using The Information In The Table Below Enter The Chegg In this case, enter the bases from top to bottom. then, for each set of three dna and complementary mrna nucleotides, use the amino acid chart to translate the nucleotides into amino acids, and type them below. enter the full, unabbreviated name of the amino acid in the blank provided. in the case of a stop codon, enter "stop". The c terminal amino acid can be determined by addition of carboxypeptidases, enzymes which cleave amino acids from the c terminal. a time course must be done to see which amino acid is released first. n terminal analysis can also be done as part of sequencing the entire protein as discussed below (edman degradation reaction). Table 5.1 the measured masses of the fragments from our example peptide. the differences in masses between the fragment masses identifies the amino acid that differentiates the two fragments. the difference in mass between fragments is used to identify the amino acid, using the following list of amino acid masses (figure 5.10). The scientist determined the number of amino acids in each of the polypeptides produced. he also counted the number of polypeptides of each length. the table below shows some of the scientist's results. use the information in the table above to calculate the number of polypeptides: 6 amino acids in length 20 amino acids in length.
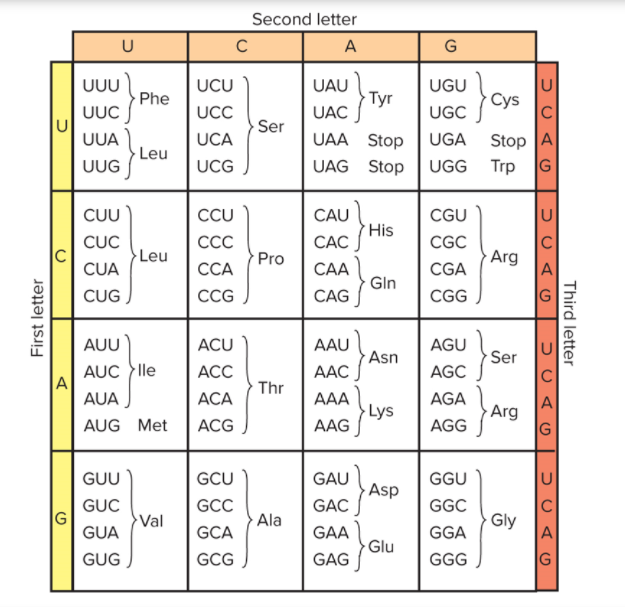
Solved The Table Shows The Amino Acids Corresponding To Chegg Table 5.1 the measured masses of the fragments from our example peptide. the differences in masses between the fragment masses identifies the amino acid that differentiates the two fragments. the difference in mass between fragments is used to identify the amino acid, using the following list of amino acid masses (figure 5.10). The scientist determined the number of amino acids in each of the polypeptides produced. he also counted the number of polypeptides of each length. the table below shows some of the scientist's results. use the information in the table above to calculate the number of polypeptides: 6 amino acids in length 20 amino acids in length. 7. 8. functional protein appears. last step. an information storage molecule called instructs a cell how to make , which do the cell's work. dna; proteins. during gene expression, a segment of is transcribed into a single stranded nucleic acid called . The central dogma: dna encodes rna; rna encodes protein. the flow of genetic information in cells from dna to mrna to protein is described by the central dogma (figure 15.3), which states that genes specify the sequence of mrnas, which in turn specify the sequence of amino acids making up all proteins.

Solved Using The Information In The Table Below Enter The Chegg 7. 8. functional protein appears. last step. an information storage molecule called instructs a cell how to make , which do the cell's work. dna; proteins. during gene expression, a segment of is transcribed into a single stranded nucleic acid called . The central dogma: dna encodes rna; rna encodes protein. the flow of genetic information in cells from dna to mrna to protein is described by the central dogma (figure 15.3), which states that genes specify the sequence of mrnas, which in turn specify the sequence of amino acids making up all proteins.

Solved Use The Amino Acid Table Below To Answer The Chegg